Furutr‰d DNA tools. Peter Larsen 2014.
Given the do-it-yourself nature of the intended user, the tool is presented with minimal documentation and without verbose explanation of analysis results.
Also note that the analysis performed by this utility is based on my own methodology and implementation, and there may be differences in the results when compared to other utilities out there.
Supports FTDNA exports only. Perhaps I will add support for other testing services exports too.
Download version 1.4c (for Windows, requires .NET framework 2.0 to run.) [200 kb]:
InstallFurutrad14c.zip
The chromosome visualization is basically a 4-way approach:
- Divide the chromosome into 4 strings of A, C, G and T's of equal length.
- Fold the strings like a 2D square using Hilbert space-filling curve algorithm.
- Apply various imageprocessing filters and pattern recognition algorithms on the squares.
- Apply colors to the squares using different presets.
The patterns themselves does not tell you anything, they are meaningless.
The purpose is to find a way of comparing and finding matching patterns between autosomal kits, but the patterns is more like admixture finding than matching autosomal segments.
Screenshot:
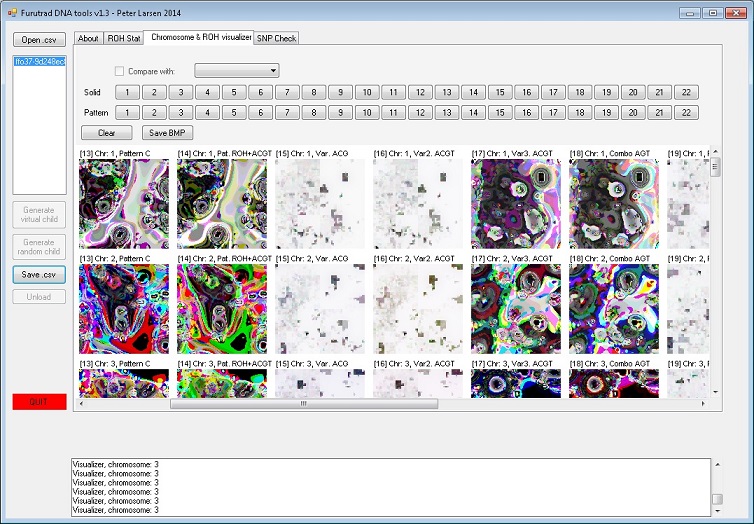
Version history:
1.4c - 2014-03-08 Added a progressbar, some bug fixes.
1.4 - 2014-03-07 Added all chromosomes in one picture visulization option
1.3 - 2014-02-17 Added Heterozygous Sequences counter
1.2 - 2014-02-17 Fixed some bugs
1.1 - 2014-02-16 Added possibility to compare two autosomal kits
1.0 - 2014-01-31 Nothing special
